Andreas Muller & Sarah Guido - Introduction to Machine Learning with Python - Chapter 2.5
Ensembles of Decision Trees
Ensembes are methods that combine multiple machine learning models.
Random Forests
The main drawback of decision trees is that they overfit the training data without pruning. A Random forest model is a collection of decision trees where each tree is slightly different from the others. The idea is that each tree might do a relatively good job at predicting but overfit in different ways, so averaging their result can reduce the overfitting while retaining the predictive power of trees.
To implement the strategy, we build many decision trees where each tree should do an acceptable job of predicting the target and should be different from each other. First we control the number of trees to build using parameter n_estimators (larger is always better!). For each tree, we randomize the selection of data points used to build a tree by taking a bootstrap sample of our data (repeatedly draw sample from data with replacement until the size is equal to the original size). Then, in each node of a tree, the algorithm randomize the selection of features used in each split test by randomly selecting a subset of features and looking for the best possible test involving one of these features. We control the size of subset using parameter max_feature. A low max_feature implies lesser similarity between each tree in the forest, less risk to overfit, and greater tree depth to fit the data well. A rule of thumb is to use max_features=sqrt(n_features) for cliassifcation and max_features=n_features for regression. Notice that in the latter, we do not randomize feature selection. Pre-pruning can also be applied using max_depth.
To make the decision in regression, every tree makes their own decision first, and their average result is reported. For calssification, each tree will provide the probability for each output class, and such probability is averaged and the class with the highest overall average probability is reported.
from sklearn.ensemble import RandomForestClassifier
from sklearn.datasets import make_moons
from sklearn.model_selection import train_test_split
X, y = make_moons(n_samples = 100, noise=0.25, random_state=3)
X_train, X_test, y_train, y_test = train_test_split(X, y, stratify=y, random_state=42)
forest = RandomForestClassifier(n_estimators=5, random_state=2)
forest.fit(X_train, y_train)
print(f"Accuracy on training set: {forest.score(X_train, y_train)}")
print(f"Accuracy on test set: {forest.score(X_test, y_test)}")
Accuracy on training set: 0.96
Accuracy on test set: 0.92
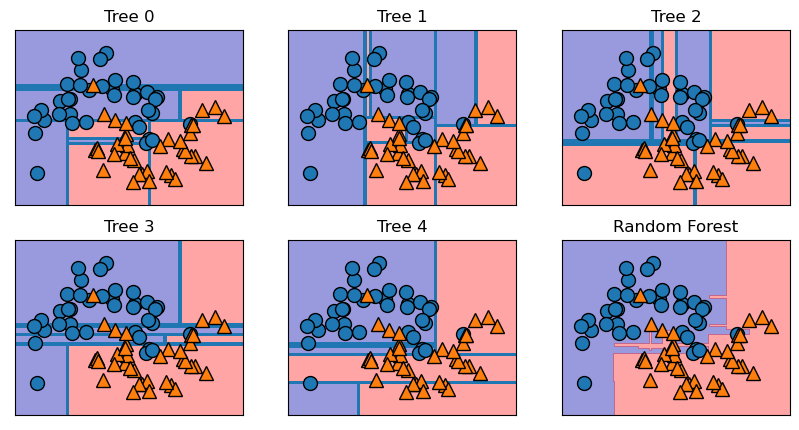
Trees built are stored in the estimator attribute. We can visualize the decision boundaries formed by each tree and their aggregated prediction. Notice each of the five trees are quite different and make some mistakes due to bootstrap sampling, while the forest overfits less and provide a more intuitive decision boundary. As more trees are added, the decision boundary will get even smoother.
import matplotlib.pyplot as plt
import mglearn
fig, axes = plt.subplots(2, 3, figsize=(10, 5))
for i, (ax, tree) in enumerate(zip(axes.ravel(), forest.estimators_)):
ax.set_title("Tree {}".format(i))
mglearn.plots.plot_tree_partition(X_train, y_train, tree, ax=ax)
mglearn.plots.plot_2d_separator(forest, X_train, fill=True, ax=axes[-1, -1], alpha=.4)
axes[-1, -1].set_title("Random Forest")
mglearn.discrete_scatter(X_train[:, 0], X_train[:, 1], y_train)
Feature importance can be assessed. If assessed, we can se ethat the model gives non-zero importance to many more features than a single tree, implying that the random forest captures a much broader picture of the data than a single tree. To sum, a random forest inherent nearly all benefits of a tree model while mitigating to some extent the overfitting.
Drawbacks of random forest include:
- Random forest is impossible to intrepret in detail given the number of trees involved
- Building trees at scale can be computationally intensive (consider using
n_jobsto control the cores used for prediction) - The forest is random. However, note that the more trees there are in the forest, the more robust the model will be against the randomness (Law of Large Number)
- Random forests do not perform wll on very high dimensional sparse data such as text data (in that case use linear models)
Gradient Boosted Regression Trees
Gradient boosting regression trees can be used both for regression and classification. It builds trees in a serial manner, where each tree tries to correct the mistakes of the previous one. Strong pre-prunning is used to achieve the process (depth of each tree is usually 1 to 5). The intuitive idea is that each tree can provide good predictions on part of the data, so more and more trees added iteratively can improve the overall performance.
Gradient boosted trees are more sensitive to parameter settings than random forests but can provide better accuracy if set correctly. Besides pre-pruning, an important parameter is the learning_rate. It controls how strongly each tree tries to correct the mistakes of the previous trees: higher learning rate implies strongr corrections and more complex models. Parameter n_estimators controls number of trees in the ensemble. A higher number of trees implies more chances to correct mistakes on the training set and higher model complexity.A common practice is to first fit n_estimatorsdepending on the time and memory budget, and then search over different learning_rates. Pre-pruning of each tree can also be applied using max_depth or max_leaf_nodes (usually max_depth is set very low).
from sklearn.datasets import load_breast_cancer
from sklearn.ensemble import GradientBoostingClassifier
cancer = load_breast_cancer()
X_train, X_test, y_train, y_test = train_test_split(cancer.data, cancer.target, random_state=0)
gbrt = GradientBoostingClassifier(learning_rate=0.01, random_state=0) #Also try max_depth=1
gbrt.fit(X_train, y_train)
print(f"Accuracy on training set: {gbrt.score(X_train, y_train)}")
print(f"Accuracy on test set: {gbrt.score(X_test, y_test)}")
Accuracy on training set: 0.9882629107981221
Accuracy on test set: 0.965034965034965
Feature importance can also be assessed for gradient boosted trees. If assessed, we can see that they also assign non-zero importance for more features, though the gradient boosted trees completely ignore some features in compared to random forests. A standard approach is to try random forests first for its robustness. But if prediction time is at a premium or we want to squzze out the last percentage of accuracy from the model, we switch to gradient boosting models. I may also look into xgboost later, which works well on large-scale datasets.
Drawbacks of gradient boosted trees include:
- Like random forests, gradient boosted trees do not work well on high-dimensional sparse data
- Gradient boosted trees also require careful tuning of parameters, as the important parameters,
n_estimatorsandlearning_rate, are closely related (e.g. lower learning tree means more trees needed to build a model of similar complexity)